![]() The primary mission of the DNA Services Lab is to provide outstanding service and state-of-the-art instrumentation to support basic, environmental, agricultural, and translational medical research for our campus investigators and external academic and corporate collaborators.
The primary mission of the DNA Services Lab is to provide outstanding service and state-of-the-art instrumentation to support basic, environmental, agricultural, and translational medical research for our campus investigators and external academic and corporate collaborators.
Instruments
Applications
Instrumentation
*We are an Illumina Certified Sequence Provider
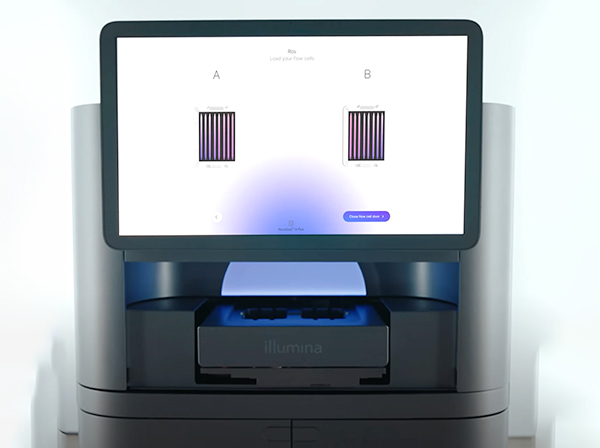

Illumina NovaSeq X Plus and 6000
The NovaSeq X Plus is the latest instrument from Illumina for ultra-high throughput sequencing of genomic DNA, RNA-Seq, DNA methylation, targeted sequencing, and many other samples that need high sequencing depth at the lowest possible cost.
The NovaSeq 6000 performs 2x250nt runs, which are still not available in the NovaSeq X Plus.
NovaSeq X Plus and 6000 Sequencing Options (see full list with multi-lane discounts in Pricing link above)
| Flowcell | Read Type and Length | Price Per Lane |
# of Single-Reads or Read Pairs Per lane |
Total # of Reads | Gb/lane |
|---|---|---|---|---|---|
| X Plus 10B | Single-reads,100bp | $1,690 | 1 billion | 1 billion | 150+ |
| X Plus 10B | Paired-reads, 2x150nt | $2,190 | 1 billion | 2 billion | 300+ |
| X Plus 25B | Paired-reads, 2x150nt | $3,290 | 3.2 billion | 6.4 billion | 900+ |
| 6000 SP | Paired-reads, 2x250nt | $3,440 | 400 million | 800 million | 200+ |

PacBio REVIO
This is the latest instrument from PacBio, it produces the longest reads with the highest accuracy versus previous PacBio instruments. It is the instrument of choice for assembly of complex genomes, metagenomes, genome resequencing with long reads, full-length 16S and other amplicons and full-length transcriptomes (IsoSeq).
One REVIO Cell produces
- up to 120 Gbases of HiFi error-corrected reads with fragments 10-30kb
- up to 13 million error corrected single-long reads.
- up to 100 million concatenated 16S, full-length cDNA or amplicon reads.
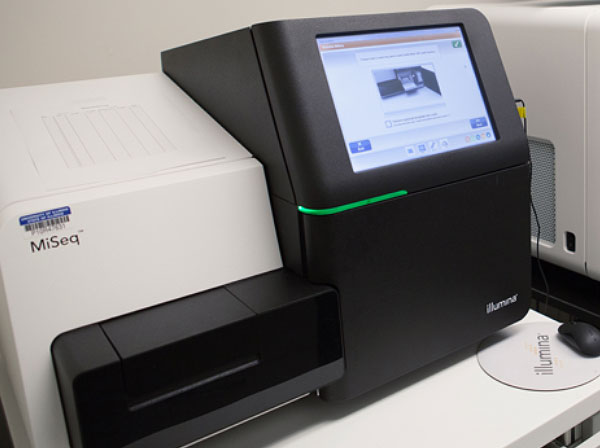

Illlumina MiSeq i100 and MiSeq
For sequencing libraries from amplicons (ie: 16S, 18S, ITS, etc), genomic DNA, microbial DNA, etc.
- The new MiSeq i100 has two kits, the 5M (5 million read-pairs) and the 25M (25 million read-pairs).
- Read length of the MiSeq i100 are up to 300bp from each side of the DNA fragments. Quality scores are very high from start to finish.
- MiSeq = V2 Nano run produces 500k to 2 million paired-reads
- MiSeq = V2 bulk run produces 8 to 20 million paired-reads
- MiSeq = V3 run produces 10 to 40 million paired-reads
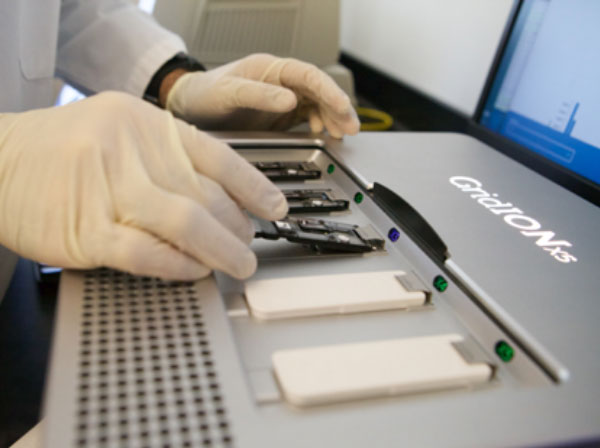

Oxford Nanopore GridION
Long sequencing reads from genomic DNA, RNA, cDNA, 16S or other amplicons.
- One flowcell produces from 1 to 9 million reads depending on the type of sample
- Reads are typically from 5kb to 40kb and with some reads from HMW genomic DNA reaching up to 100kb
- Each flowcell runs one or multiple samples
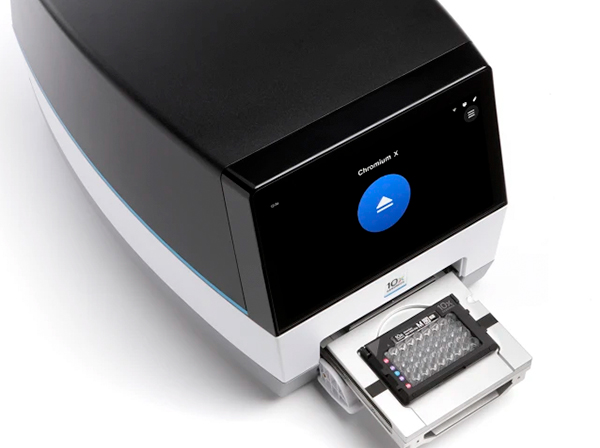

10x Genomics
The 10x iX Chromium system produces libraries for 3' transcriptome sequencing from single-cells, ATAC-seq and multiomics (RNAseq + ATACseq).
- Libraries constructed on 10x are fully compatible with Illumina sequencing.
- Single-cell libraries can be constructed from 500 to 10,000 cells per library.
- Single-cell libraries are sequenced for 28x100nt or 28x150nt on the NovaSeq, with a minimum of 50k reads per cell recommended.
- 10x Visium Spatial gene expression libraries are also available.
- 10x Xenium in situ platform is also available in our sister Core, CMtO.